
Producing the ENSO index verification heatmaps#
In this example we use the computed El Niño–Southern Oscillation (ENSO) indices from the previous Notebook to produce heatmaps of temporal correlations, as included on the C3S seasonal real-time forecast verification page.
Some information on ENSO impacts in Europe can be found on a page in the C3S documentation.
Load the pre-computed ERA5 indices and prepare to compute correlations
Loop over the hindcast start months and compute correlations
Note
Some of the python code uses a set of utility functions available in the GitHub respository which hosts these Notebooks.
Configuration#
Import required modules, define the library of C3S systems, and define some parameters related to the indices previously computed.
import os
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import xarray as xr
import regionmask
import seaborn as sns
from matplotlib.colors import BoundaryNorm
from dateutil import relativedelta
import sys
from pathlib import Path
# Add parent directory to path so we can import the utils package
sys.path.insert(0, str(Path.cwd().parent))
from utils.tools import getForecastsystemDetails
# the data path where the indices were saved
data_path = 'data'
# define the common hindcast period
hc_period = ['1993', '2016']
hc_str = '_'.join([hc_period[0], hc_period[1]])
# select a forecast system
origin = 'ukmo' # originating centre label for the CDS request
system = '604' # forecast system identifier for the CDS request
prov = f'{origin}.s{system}'
# Get system details from configuration files in utils directory using format ukmo.s604
system_details = getForecastsystemDetails(prov)
print(system_details)
{'origin_details': {'MARS_origin': 'egrr', 'label': 'Met Office'}, 'system_details': {'name': 'GloSea6 (system=604)', 'isLagged': True, 'ens_size': {'products': {'fc': 50}}}}
Load the pre-computed ERA5 indices and prepare to compute correlations#
# load in reanalysis
era5_file_name = '/era_5_nino_ind_1993_2016.nc'
era5_ind = xr.open_dataarray(data_path + era5_file_name)
era5_ind
# create a correlation dataset
corr = xr.Dataset()
Loop over the hindcast start months and compute correlations#
Loop over the start months. For each start month do the following:
deal with the missing Jan 1993 start if the centre is
ukmocreate a copy of the ERA5 array (to rearrange to match the valid times of the start month in question)
read in the hindcast index for the given start, and refactor to include valid time
sub-select qualifying ensemble members, if needed (for forecast systems where not all available ensemble members are used in the graphical products)
refactor the ERA5 array to align with the hindcast leadtimes
for each leadmonth compute the temporal correlation
combine the leadmonth correlations and append to the array of correlations by start month
The result is an array of temporal correlations for each start month and leadmonth.
for stmonth in range(1, 13):
print(f'START MONTH = {stmonth}')
# Note that if the chosen system does not yet have all starts then this needs to be dealt with here
# This also includes January 1993 for UKMO meaning the ERA5 data needs to be adjusted to match this
# Met Office systems missing data - start month
if origin == 'ukmo' and stmonth == 1:
# edit config to deal with Jan 1993 data gap in Met Office data
hc_period[0] = '1994'
o = era5_ind.sel(time=slice(hc_period[0], hc_period[1])).copy()
else:
o = era5_ind.copy()
# read hindcast data array for current start
hc_file_name = f'/sst_mm_{origin}_{system}_{hc_str}_st_{stmonth}_nino_ind.nc'
hcst = xr.open_dataarray(data_path + hc_file_name)
#hcst = hcst.reset_coords(names="valid_time", drop=True)
vt = xr.DataArray(dims=('start_date', 'forecastMonth'),
coords={'forecastMonth': hcst.forecastMonth, 'start_date': hcst.start_date})
vt.data = [[pd.to_datetime(std) + relativedelta.relativedelta(months=fcmonth - 1) for fcmonth in vt.forecastMonth.values] for
std
in vt.start_date.values]
hcst = hcst.assign_coords(valid_time=vt)
# select qualifying ensemble members
# Check if ensemble size in data exceeds the configured size
ens_size_hc = system_details['system_details'].get('ens_size', {}).get('products', {}).get('hc',None)
if ens_size_hc is not None and len(hcst.number) > ens_size_hc:
hcst = hcst.sel(number=slice(None, ens_size_hc - 1))
# refactor reanalysis to align with hindcast leadtimes
o = o.rename({'time': 'start_date'}).swap_dims({'start_date': 'valid_time'})
fcmonths = [mm + 1 if mm >= 0 else mm + 13 for mm in
[t.month - stmonth for t in pd.to_datetime(o.valid_time.values)]]
o = o.assign_coords(forecastMonth=('valid_time', fcmonths))
o = o.loc[(o['forecastMonth'] <= 6)]
# loop over forecast month, define the hindcast mean over the ensemble members, and compute the correlation over the hindcast years
l_corr = list()
l_corr_det = list()
for this_fcmonth in hcst.forecastMonth.values:
thishcst = (hcst.sel(forecastMonth=this_fcmonth).swap_dims({'start_date': 'valid_time'}))
thishcst = thishcst.mean('number')
thisobs = o.where(o.valid_time == thishcst.valid_time, drop=True)
l_corr.append(xr.corr(thishcst, thisobs, dim='valid_time'))
thiscorr = xr.concat(l_corr, dim='forecastMonth')
thiscorr = thiscorr.assign_coords(forecastMonth=[1, 2, 3, 4, 5, 6])
thiscorr = thiscorr.expand_dims(dim={'start_month': [stmonth]}, axis=0)
corr = xr.merge([corr, thiscorr], compat='no_conflicts', join='outer')
Save the result, and inspect it.
file_name = f'/sst_mm_{origin}_{system}_{hc_str}_nino_ind_corr.nc'
corr.to_netcdf(data_path + file_name, mode='w')
corr
<xarray.Dataset> Size: 3kB
Dimensions: (start_month: 12, region: 4, forecastMonth: 6)
Coordinates:
* start_month (start_month) int64 96B 1 2 3 4 5 6 7 8 9 10 11 12
* region (region) int64 32B 0 1 2 3
* forecastMonth (forecastMonth) int64 48B 1 2 3 4 5 6
surface float64 8B 0.0
abbrevs (region) <U3 48B '1+2' '3' '3.4' '4'
names (region) <U7 112B 'NINO1+2' 'NINO3' 'NINO3.4' 'NINO4'
number int64 8B 0
step timedelta64[ns] 8B 00:00:00
Data variables:
sst (start_month, forecastMonth, region) float64 2kB 0.9424 .....Plot the correlation heatmaps#
Using the computed correlations, construct correlation heatmaps for each NINO index computed in the previous Notebook.
Define labels (valid months or start months, origin and system) and load the correlations.
import calendar
mon_labels = [m.lower() for m in calendar.month_abbr[1:]]
orig_label = system_details.get('origin_details', {}).get('label', 'UnknownOrigin')
sys_name = system_details.get('system_details',{}).get('name','UnknownSystem')
# load the correlations
file_name = f'/sst_mm_{origin}_{system}_{hc_str}_nino_ind_corr.nc'
corr = xr.open_dataarray(data_path + file_name)
Loop over the regions for which the indices and correlations were computed:
refactor the correlations for each region/index in a pandas dataframe (remove unnecessary dimensions and variables)
plot correlation as a function of start month and leadtime using a seaborn heatmap
refactor the correlations again to include valid month instead of start month
plot correlation as a function of valid month and leadtime
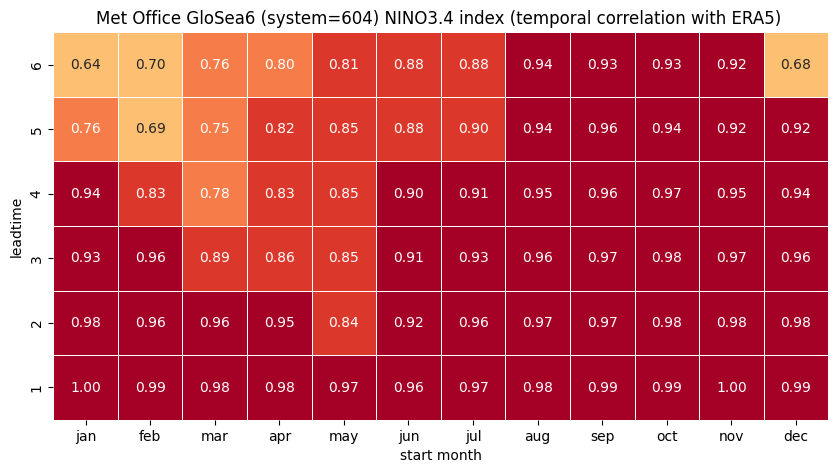
# make the plot - loop over nino regions
for ind in corr.region.values:
print(ind, corr.sel(region=ind).names.values)
ind_name = str(corr.sel(region=ind).names.values)
hm_cm = plt.get_cmap('RdYlBu_r')
hm_levs = np.linspace(0, 1, 11)
norm = BoundaryNorm(hm_levs, ncolors=hm_cm.N, clip=True)
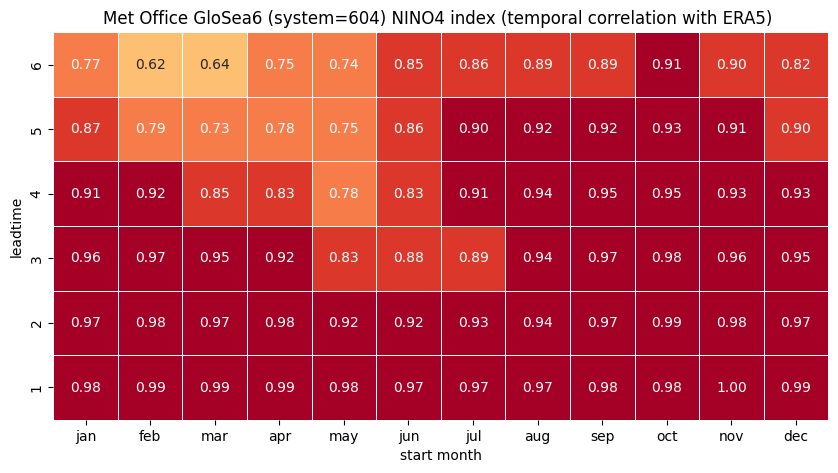
# HEATMAP (correlation as a function of start_month and leadtime)
df = (corr.sel(region=ind).to_dataframe()
.drop(['region', 'surface', 'abbrevs', 'step', 'names', 'number'], axis=1))
df = df.unstack().droplevel(0, axis=1)
fig = plt.figure(figsize=(10, 10))
ax = sns.heatmap(df.T, square=True, cmap=hm_cm, vmin=hm_levs.min(), vmax=hm_levs.max(), norm=norm,
annot=True, fmt=".2f", cbar=False, linewidths=0.5, linecolor='white')
ax.grid(False)
ax.invert_yaxis()
tit_txt1 = f'{ind_name} index (temporal correlation with ERA5) '
tit_txt2 = f'{orig_label} {sys_name} '
plt.title(tit_txt2 + tit_txt1)
plt.xlabel('start month')
plt.ylabel('leadtime')
# Set ticks labels for x-axis
ax.set_xticklabels(mon_labels)
ax.set_yticklabels(['1', '2', '3', '4', '5', '6'])
plt.show()
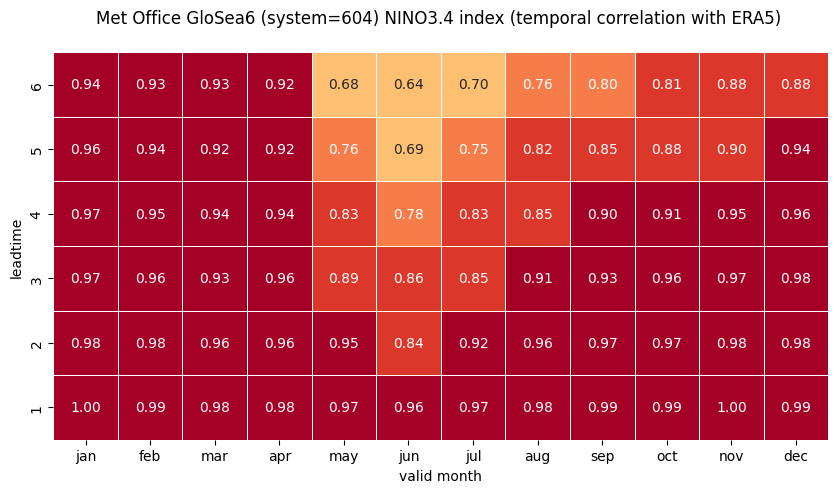
# HEATMAP (correlation as a function of valid_month and leadtime)
df_vt = df.stack(future_stack=True).reset_index()
df_vt['valid_month'] = df_vt.apply(lambda row: row['start_month']
+ row['forecastMonth'] - 1, axis=1).astype('int32')
df_vt['valid_month'] = [int(mm) if mm < 13 else mm - 12 for mm in df_vt['valid_month']]
df_vt = df_vt.drop('start_month', axis=1).set_index(
['valid_month', 'forecastMonth']).unstack().transpose().droplevel(0)
fig = plt.figure(figsize=(10, 10))
ax = sns.heatmap(df_vt, square=True, cmap=hm_cm, vmin=hm_levs.min(), vmax=hm_levs.max(), norm=norm,
annot=True, fmt=".2f", cbar=False, linewidths=0.5, linecolor='white')
ax.grid(False)
ax.invert_yaxis()
plt.title('\n'.join([tit_txt2 + tit_txt1, '']))
plt.xlabel('valid month')
plt.ylabel('leadtime')
# Set ticks labels for x-axis
ax.set_xticklabels(mon_labels) # , rotation='vertical', fontsize=18)
plt.show()
0 NINO1+2
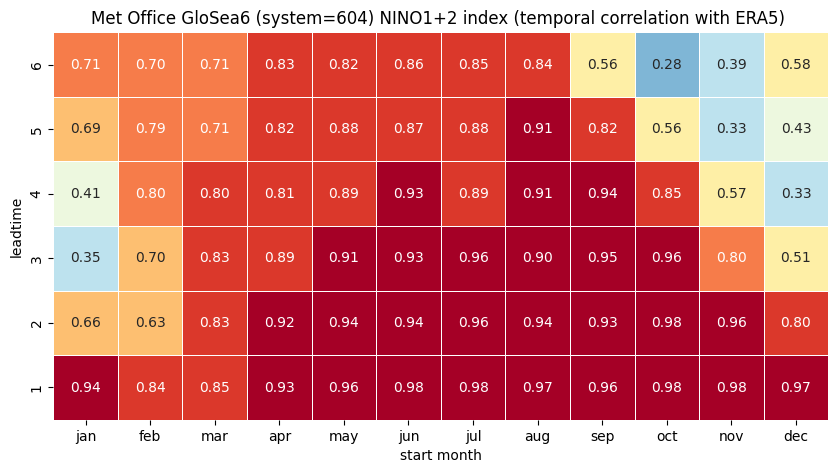
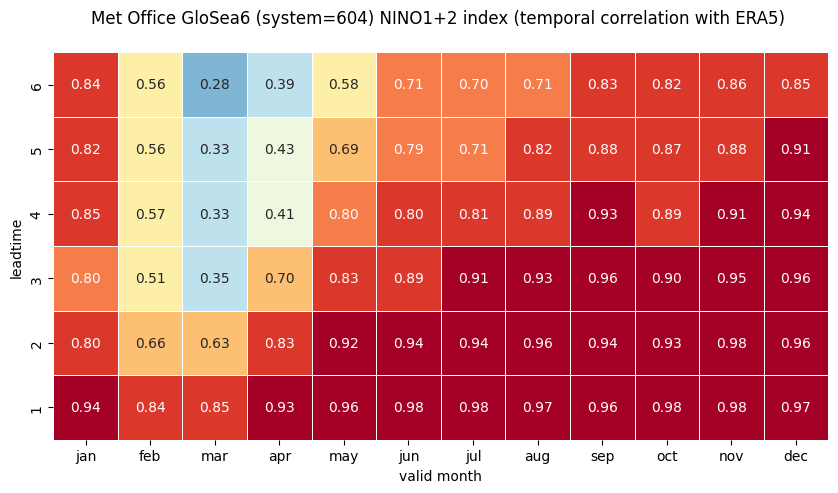
1 NINO3
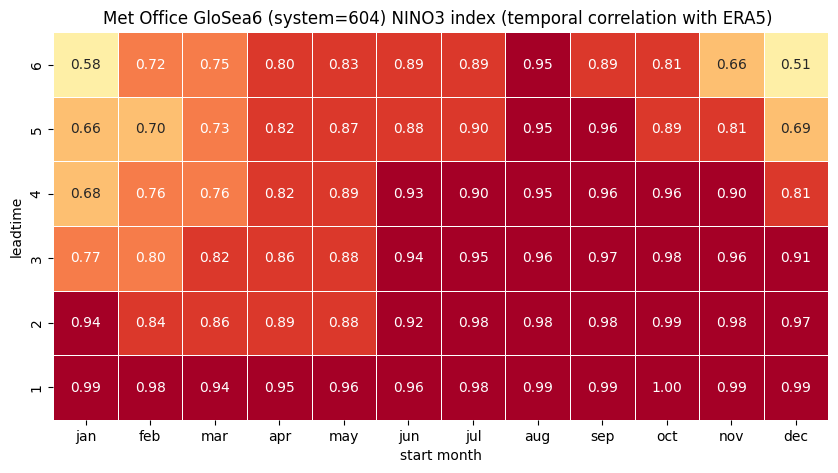
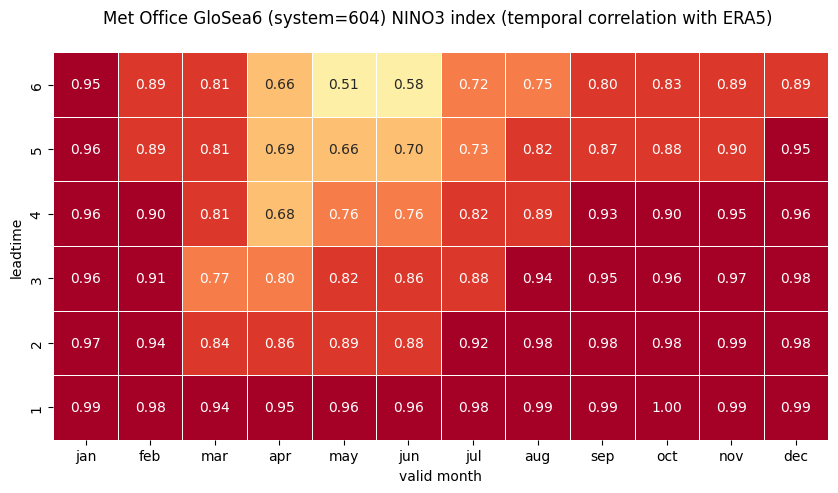
2 NINO3.4
3 NINO4
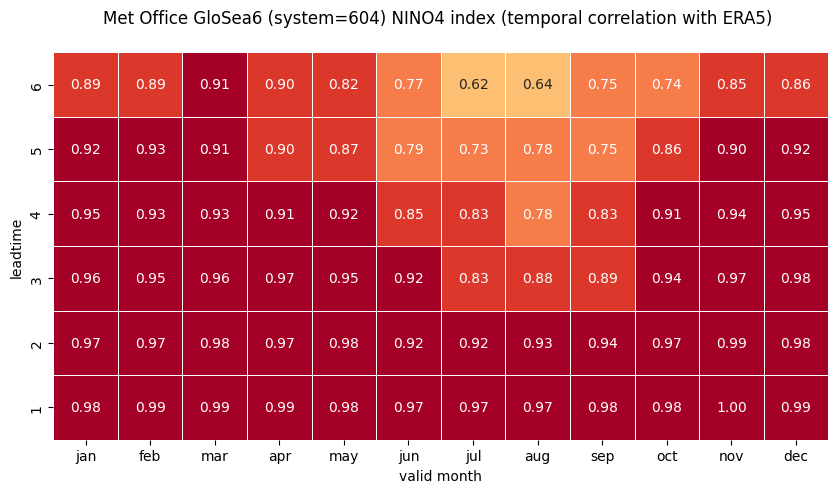
Extensions to these single-system monthly correlations heatmaps would be to compute indices for multiple systems and then combine them into multi-system combinations. The approach to variance-normalisation in the C3S SST index charts is outlined on the products description page, and in the SEAS5 user guide.
